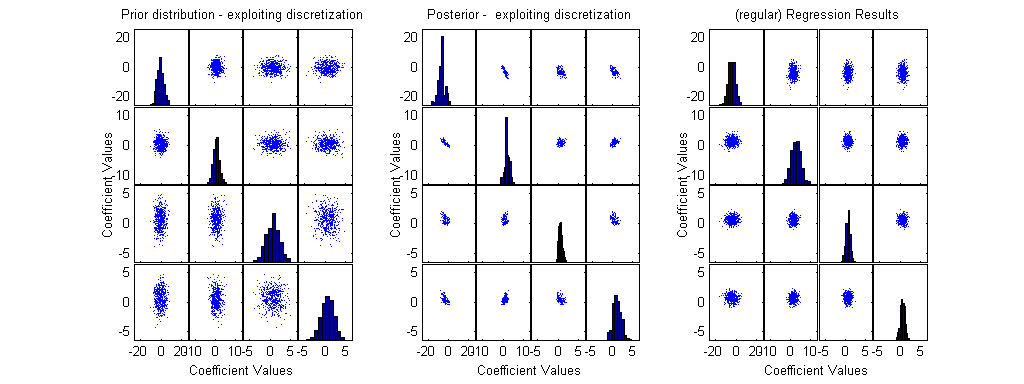
We suggst a method which allows estimation of posterior information even when the closed form of the posterior is very complex, exploiting a discretization of the prior distribution.
This need for posterior distribution is quite frequent, and we use it for the purpose of analyzing small sample data with generalized linear models. Using regression techniques for GLM with small samples is not reliable, and often highly biased.
The sequential experimental design examples already contain this utilization; here this idea is given a seperate example, for ready-made data.
This file contains a Poisson example. X0 contains the location of the experiment, RResult contains the responses observed.
We provide 3 plots: (1) A presentation of the prior, by a sample of possible discrete points; (2) A presentation of the posterior, by discrete points; (3) A sample of possible coefficient values, from analysis through regression (GLM).
This script was written by Hovav Dror, Tel-Aviv University, 2006
If you make changes to the script, or if you implement it to a (to be) published problem, please inform us: dms@post.tau.ac.il ; hovavdror@gmail.com
further details, including papers describing the algorithm, can be found at: http://www.math.tau.ac.il/~dms/GLM_Design
NN=10000; % Number of parameter vectors to represent the prior using a discretization m=3; p=4; BetaMeans=[ -0.7 0.5 0.7 0.7]; BetaSE=[3 1.5 1.5 1.5]; NI=rand(NN,p); beta = norminv(NI,ones(NN,1)*BetaMeans,ones(NN,1)*BetaSE); family='poisson'; link='log'; model='linear';
Locations
X0=[
0 0 2
2 2 0
2 0 2
2 2 2
1 3 3
];
F0=x2fx(X0,model);
% Results:
RResult=[
0
1
1
3
3
];
scrsz = get(0,'ScreenSize'); set(gcf,'Position',[0 scrsz(4) scrsz(3) scrsz(4)/2]); % small figure upper right of the screen subplot(1,3,1); [H,AX,BigAx,P] =plotmatrix(beta(1:500,:)); % A sample of 500 points from the prior xlabel('Coefficient Values'); ylabel('Coefficient Values'); title('Prior distribution - exploiting discretization'); x1=get(AX,'XLim'); y1=get(AX,'YLim');
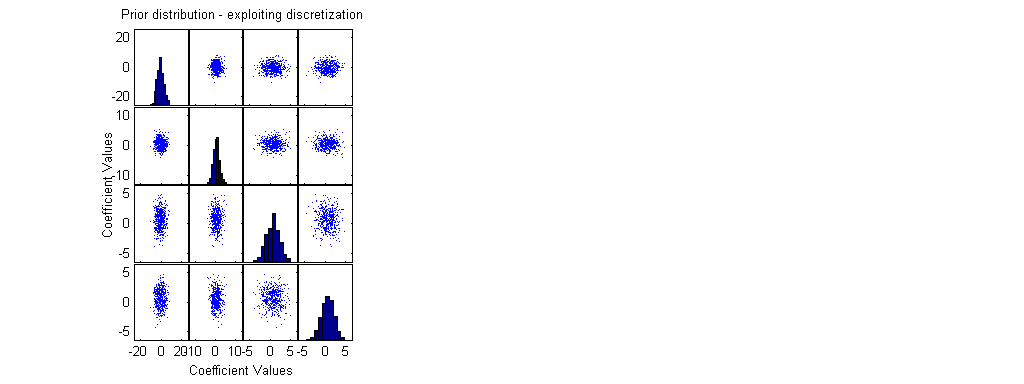
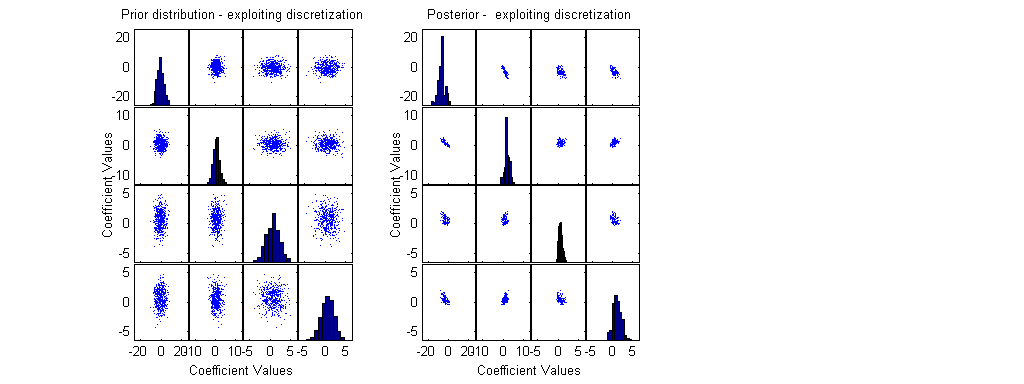
ResultFactorial=[]; for i=1:length(RResult) if RResult(i)<21 ResultFactorial(end+1)=factorial(RResult(i)); else ResultFactorial(end+1)=-1; % else - use normal approximation and not the exact poisson distribution end; end; RealMinSS=realmin^(1/length(RResult)); RResultM=ones(NN,1)*RResult'; bx=(F0*beta')'; lambda=exp(bx); LRi=[]; for i=1:NN LRi(i)=0; for j=1:length(RResult) if lambda(i,j)>20 | RResult(j)>20 % if it is better to use normal approximation LRi(i)=LRi(i) -0.5*log(2*pi*lambda(i,j)) - (RResult(j) - lambda(i,j))^2 / (2*lambda(i,j)) ; else % for small lambda & RResult use exact poisson distribution LRi(i)=LRi(i) +log( max([lambda(i,j)^RResult(j) * exp(-lambda(i,j)) / ResultFactorial(j) RealMinSS])); end; % if lambda end; % for j end; % for i MaxLogLike=max(LRi); MinLogLike=min(LRi); LRealMin=log(realmin); if MinLogLike<LRealMin ScaleAddition=LRealMin-(MinLogLike-MaxLogLike); LRi=LRi+ScaleAddition; MaxLogLike=max(LRi); end; if exp(MaxLogLike)>0 % if max likelihood is computable LRi1=exp(LRi-MaxLogLike); LRi1=LRi1/sum(LRi1); else fprintf('\nMax Likelihood =0 ! \n'); end; Weights=LRi1';
Wbeta=sortrows([Weights beta]); SW=cumsum(Wbeta(:,1)); beta1=Wbeta(SW>0.05,2:end); % Posterior possible beta values, for a 95% level Weights1=Wbeta(SW>0.05,1); Sample1=randsample(length(Weights1),500,true,Weights1); % Choose a Weighted sample of size 500 subplot(1,3,2); [H2,AX2,BigAx2,P2] =plotmatrix(beta1(Sample1,:)); for i=1:p for j=1:p set(AX2(j,i),'XLim',x1{(i-1)*p+1}); set(AX2(i,j),'YLim',y1{i}); end; end; title('Posterior - exploiting discretization'); xlabel('Coefficient Values'); ylabel('Coefficient Values');
subplot(1,3,3); [b,dev,stats] = glmfit(X0,RResult,'poisson'); NI2=rand(500,p); beta2 = norminv(NI2,ones(500,1)*b',ones(500,1)*stats.se'); [H3,AX3,BigAx3,P3] =plotmatrix(beta2); for i=1:p for j=1:p set(AX3(j,i),'XLim',x1{(i-1)*p+1}); set(AX3(i,j),'YLim',y1{i}); end; end; title('(regular) Regression Results'); xlabel('Coefficient Values'); ylabel('Coefficient Values');